IDEA CAPSULES (SMALL UNITS OF INFORMATION PACKED INTO BUNDLES. PERFECT FOR REVISION)
IDEA CAPSULE 1- Intro to Genetic Variation
The achievment standard 91157, "Demonstrate understanding of genetic variation and change" covers several key ideas relating to genetic variation which are interlinked.
I generally like to divide this achievement standard into two "sections", as I believe that these are two overarching themes within this standard.
The achievment standard 91157, "Demonstrate understanding of genetic variation and change" covers several key ideas relating to genetic variation which are interlinked.
I generally like to divide this achievement standard into two "sections", as I believe that these are two overarching themes within this standard.
- Section 1- Genetic Variation in Individuals and Inheritance. This section begins with a look at meiosis (i.e. how sex cells divide to produce gametes). We will look at the reasons why the process of meiosis generates so much genetic variation in offspring, hence why we don't look exactly the same as our siblings (unless you are an identical twin!). We'll follow on from this, looking at inheritance and how different traits are inherited by offspring. We will be using Punnett squares to determine problems involving both monohybrid inheritance (inheritance of a single trait) and dihybrid inheritance (inheritance of two different traits).
- Section 2- Genetic Variation within a Population. This section covers how allele frequencies vary in populations. Some of the main topics include: How movement, population size, random/non-random mating, and selection can have an impact on the allele frequencies within populations. We will also look at different types of selection (sexual, natural and artificial) as well as look at two occurences which can also affect allele frequencies within populations (Founder's Effect and the Bottleneck Effect).
BEFORE WE START: Genetic Variation Vocab Checklist
Download and use this Genetic Variation Vocab Checklist to familiarise yourself with some of the key words we will be using in this unit. When we come across these words in class- write a definition in your own words in the box beside the word to complete your own vocabulary chart. Your aim is to be able to use these words confidently and correctly in your writing when referring to different contexts.
Download and use this Genetic Variation Vocab Checklist to familiarise yourself with some of the key words we will be using in this unit. When we come across these words in class- write a definition in your own words in the box beside the word to complete your own vocabulary chart. Your aim is to be able to use these words confidently and correctly in your writing when referring to different contexts.
| Genetics Vocabulary Checklist | |
| File Size: | 13 kb |
| File Type: | docx |
IDEA CAPSULE 2- Meiosis
Process of Meiosis
Unlike mitosis (which you will recall is when diploid body/ somatic cells divide to produce two identical body/ somatic cells), meiosis occurs in sex cells where cells go through two stages of division, resulting in four different haploid gametes (one set of chromosomess per cell).
The process by which the sex cells divide in meiosis is similar to the process we have already covered in mitosis. However, there are two phases of division in meiosis instead of one.
In more detail....
The process of meiosis begins rather like mitosis, however it goes through two rounds of PMAT (Prophase, Metaphase, Anaphase, Telophase).
Use the resources below (PowerPoint, worksheet, animations and video) to understand what occurs during meiosis.
(Note: The names of the phases are not important, however they may be helpful to refer to when writing).
MEIOSIS POWERPOINT
Process of Meiosis
Unlike mitosis (which you will recall is when diploid body/ somatic cells divide to produce two identical body/ somatic cells), meiosis occurs in sex cells where cells go through two stages of division, resulting in four different haploid gametes (one set of chromosomess per cell).
The process by which the sex cells divide in meiosis is similar to the process we have already covered in mitosis. However, there are two phases of division in meiosis instead of one.
- In meiosis I, the chromosomes will condense into chromosomes.
- Then the chromosomes will begin to line up in their homologous pairs.
- After this, the homologous pairs separate and are pulled away from each other by the spindle fibres.
- Two different cells are formed. Each is haploid (i.e. has one set of chromosomes)
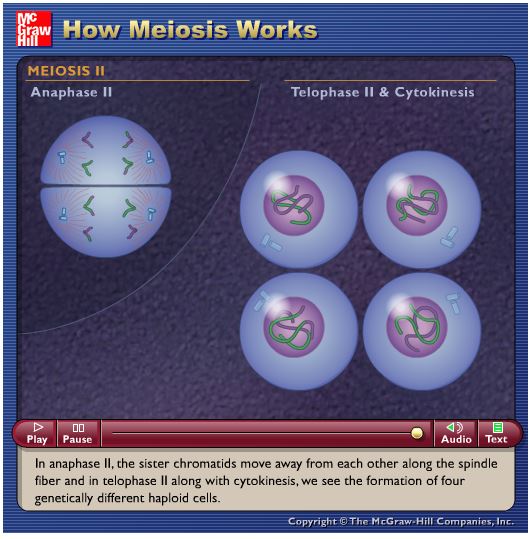
- In meiosis II, the chromosomes line up individually down the middle and the chromatids are split apart from each other by the spindle fibres.
- In total, four unique haploid ( half the amount of genetic information as the original diploid cell) cells are created. These are called gametes (i.e. eggs and sperm in humans).
In more detail....
The process of meiosis begins rather like mitosis, however it goes through two rounds of PMAT (Prophase, Metaphase, Anaphase, Telophase).
Use the resources below (PowerPoint, worksheet, animations and video) to understand what occurs during meiosis.
(Note: The names of the phases are not important, however they may be helpful to refer to when writing).
MEIOSIS POWERPOINT
MEIOSIS WORKSHEET
Use this alongside the videos and animations to jot down notes.
| Meiosis Worksheet | |
| File Size: | 53 kb |
| File Type: | docx |
MEIOSIS ANIMATIONS
|
Sumasinc- Meiosis animation and quiz
|
|
McGraw Hill- Meiosis animation and quiz
MEIOSIS VIDEOS |
|
|
|
How does meiosis contribute to genetic variation? (IMPORTANT)
We are interested in meiosis because it creates the opportunity for a lot of genetic variation, through a number of different happenings which occur throughout meiosis. The ways that meiosis contributes to genetic variation within a new organism are outlined in the bullet points below:
We are interested in meiosis because it creates the opportunity for a lot of genetic variation, through a number of different happenings which occur throughout meiosis. The ways that meiosis contributes to genetic variation within a new organism are outlined in the bullet points below:
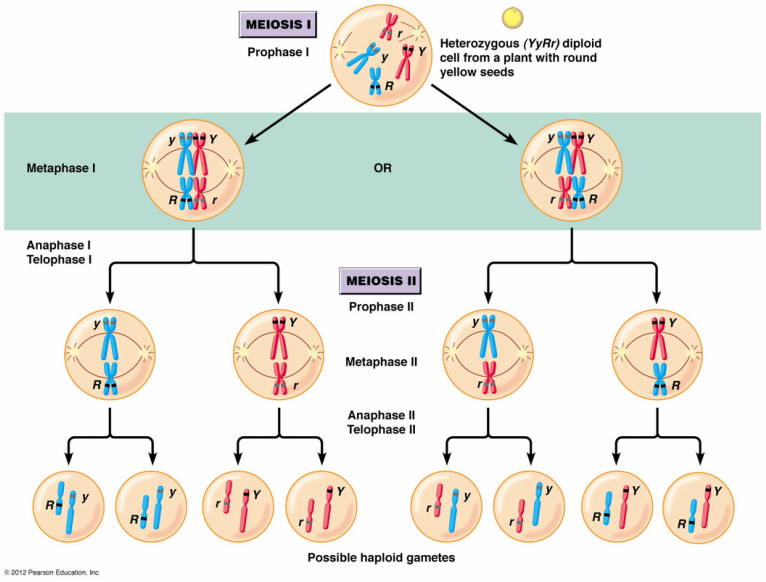
- Independent assortment- occurs when the homologous pairs of chromosomes line up in a random order during metaphase 1 of meiosis- for humans with 23 pairs of chromosomes there are more than 8 million possible combinations of chromosomes in the gametes that are produced (so much variation already!!! This is why you don’t look exactly like your sibling- unless of course you are identical twins).
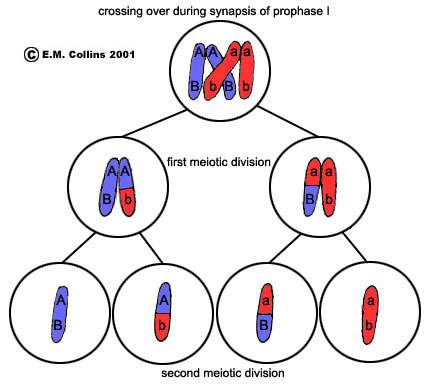
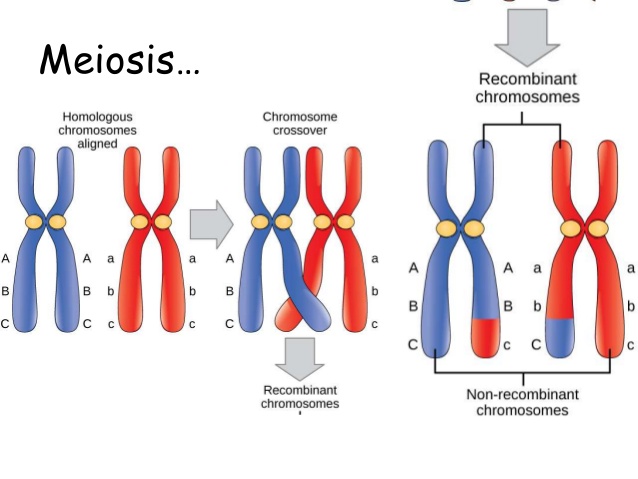
- Crossing over and recombination- Alleles that are on the same chromosome are linked because they tend to be inherited together. However, linked genes can be separated by crossing over which occurs in prophase of Meiosis 1 i.e. when homologous chromosomes are paired up.
- Segregation- the independent separation of alleles on different chromosomes (e.g. recessive or dominant allele and the combination of those that are on each chromosome). This occurs during anaphase II of meiosis when the sister chromatids separate from each other.
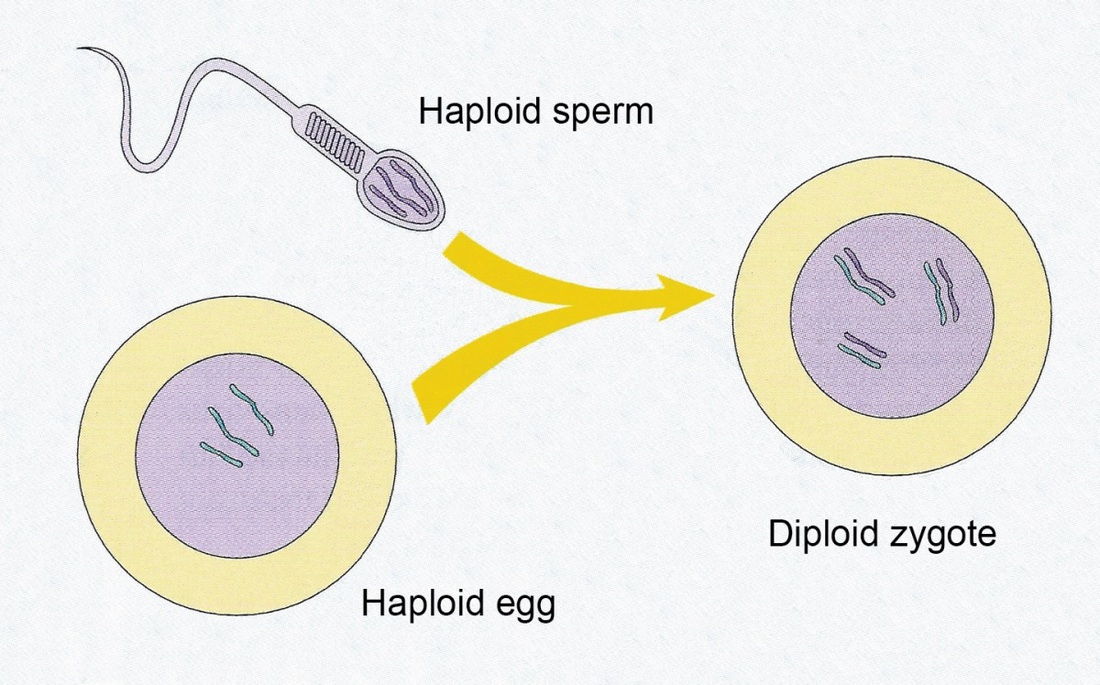
- Fertilisation- Fusion of sperm and ovum (egg) which are both haploid cells to bring together the maternal and paternal chromosomes to restore the diploid state so the offspring has a complete set of chromosomes. Such a huge amount of variation is again made possible by the fact that both the ovum and sperm have unique combinations of alleles due to independent assortment, crossing over and segregation PLUS it is up to chance which sperm fertilises the ovum, resulting in an individual with a unique set of alleles.
IDEA CAPSULE 3- Monohybrid inheritance
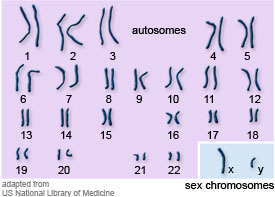
Chromosomes and Sex chromosomes- Of the 26 chromosomes in our cells, 22 exist as homologous pairs and are known as autosomes- these have genes that contribute information for all our general body structures and functions.
Then there are two sex chromosomes which determine the sex of the organism. Females are XX, Males are XY. The Y chromosome is much shorter chromosome than X hence has far fewer genes. (Look at the bottom right hand corner of this diagram for reference).
Karyotype- We often create a karyotype to study the chromosomes present in an individual. To make a human karyotype, the chromosomes are stained, matched for size and shape and then numbered and grouped into their homologous pairs from 1-22 along with the sex chromosomes. (See photo above).
What is monohybrid inheritance?
Monohybrid inheritance is the inheritance of one characteristic controlled by one gene which can have two or more different alleles at the same locus on a homologous pair of chromosomes.
Complete dominance- Complete dominance occurs when the information on one allele (the dominant allele) is always expressed in the phenotype (physical appearance) when that allele is present in the genotype (genetic makeup) à i.e. the presence of a dominant allele masks the presence of a recessive allele. A recessive allele will only be expressed in the phenotype when both alleles in the genotype are recessive e.g. aa.
When both alleles in a genotype are the same e.g. AA or aa, the organism is considered homozygous and a pure breeder because it can only pass on one type of allele to its offspring. However when both alleles are different e.g. Aa, the individual is heterozygous for the trait (and is not a pure breeder because it can pass on either type of allele to its offspring).
Monohybrid cross- A monohybrid cross is the study of the inheritance of one characteristic. You need to be able to explain a monohybrid cross in terms of genotypes. This can be done by writing down the genotype of the parents and the gametes which each can produce in a special table called a Punnett Square (we will cover this next). After you have completed your Punnett square, you can then determine both genotype and phenotype ratios for the offspring of the two parents.
Chromosomes and Sex chromosomes- Of the 26 chromosomes in our cells, 22 exist as homologous pairs and are known as autosomes- these have genes that contribute information for all our general body structures and functions.
Then there are two sex chromosomes which determine the sex of the organism. Females are XX, Males are XY. The Y chromosome is much shorter chromosome than X hence has far fewer genes. (Look at the bottom right hand corner of this diagram for reference).
- Whether a child is male or female is determined at the moment of fertilisation by the sperm because the sperm has either X or Y chromosomes due to meiosis (because they have 50% chance of receiving an X or a Y chromosome from the father).
- Eggs are always X (because they will always receive an X from the mother).
- “It is the presence or absence of Y that determines sex; an individual with a Y is a male whilst an individual without a Y is female.
Karyotype- We often create a karyotype to study the chromosomes present in an individual. To make a human karyotype, the chromosomes are stained, matched for size and shape and then numbered and grouped into their homologous pairs from 1-22 along with the sex chromosomes. (See photo above).
What is monohybrid inheritance?
Monohybrid inheritance is the inheritance of one characteristic controlled by one gene which can have two or more different alleles at the same locus on a homologous pair of chromosomes.
Complete dominance- Complete dominance occurs when the information on one allele (the dominant allele) is always expressed in the phenotype (physical appearance) when that allele is present in the genotype (genetic makeup) à i.e. the presence of a dominant allele masks the presence of a recessive allele. A recessive allele will only be expressed in the phenotype when both alleles in the genotype are recessive e.g. aa.
When both alleles in a genotype are the same e.g. AA or aa, the organism is considered homozygous and a pure breeder because it can only pass on one type of allele to its offspring. However when both alleles are different e.g. Aa, the individual is heterozygous for the trait (and is not a pure breeder because it can pass on either type of allele to its offspring).
Monohybrid cross- A monohybrid cross is the study of the inheritance of one characteristic. You need to be able to explain a monohybrid cross in terms of genotypes. This can be done by writing down the genotype of the parents and the gametes which each can produce in a special table called a Punnett Square (we will cover this next). After you have completed your Punnett square, you can then determine both genotype and phenotype ratios for the offspring of the two parents.
IDEA CAPSULE 4- Punnett Squares for monohybrid crosses
If you are new to Punnett Squares or need a little refresher, have a look at the document below. It will help you set up your own Punnett Squares for monohybrid crosses (inheritance of a single trait) so that you can work out genotype and phenotype ratios of offspring from two parents.
If you are new to Punnett Squares or need a little refresher, have a look at the document below. It will help you set up your own Punnett Squares for monohybrid crosses (inheritance of a single trait) so that you can work out genotype and phenotype ratios of offspring from two parents.
|
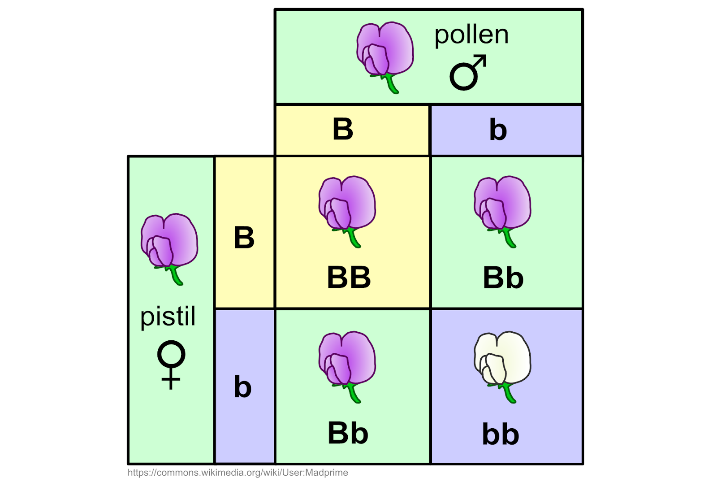
Another example of a monohybrid cross: Flower colour in pea plants Pea plants can either have purple or white flowers. The purple colour shows complete dominance over white colour. Alleles
In this cross, the Mother is heterozygous (Bb) and Father is also heterozygous (Bb) for the trait. Offspring produced have:
|
IDEA CAPSULE 5- Pedigree charts
A pedigree chart is a diagram that shows the occurrence and appearance or phenotypes of a particular gene or organism and its ancestors from one generation to the next, most commonly humans, show dogs, and race horses.
Note, that you may see the term "Pure-breeding" used when referring to pedigree charts. This means that the individual is homozygous (either recessive or dominant) for that trait. I.e. it can only pass down one type of allele to its offspring.
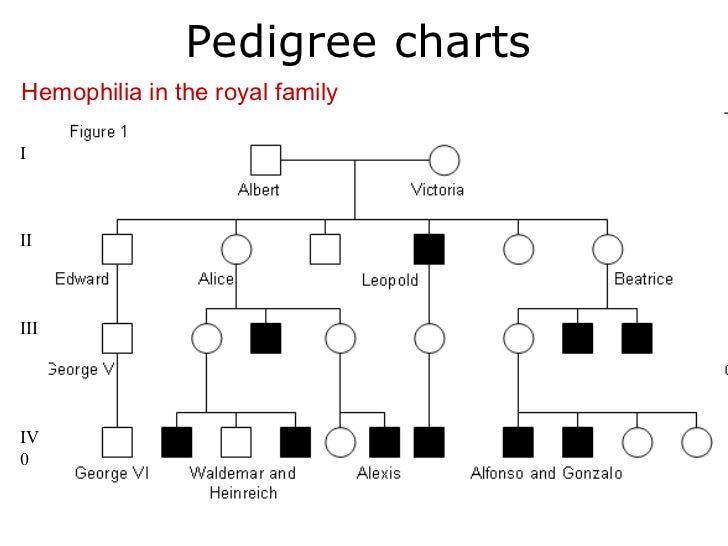
The following diagram presents a pedigree chart detailing hemophilia (a rare disorder in which the blood does not clot normally) in the Royal Family. By looking at the pedigree chart we can determine which members of the Royal Family have hemophilia (a phenotype).
A pedigree chart is a diagram that shows the occurrence and appearance or phenotypes of a particular gene or organism and its ancestors from one generation to the next, most commonly humans, show dogs, and race horses.
Note, that you may see the term "Pure-breeding" used when referring to pedigree charts. This means that the individual is homozygous (either recessive or dominant) for that trait. I.e. it can only pass down one type of allele to its offspring.
The following diagram presents a pedigree chart detailing hemophilia (a rare disorder in which the blood does not clot normally) in the Royal Family. By looking at the pedigree chart we can determine which members of the Royal Family have hemophilia (a phenotype).
- A square in this diagram represents a male
- A circle in this diagram represents a female
- Black coloured shapes represent those affected by hemophilia
- Light coloured shapes represent those unaffected by hemophilia
Application of Pedigree charts:
Some NCEA questions may require you to determine the phenotype of offspring through a pedigree chart.
We can do this by deducing parent genotypes and then working out possible offspring genotypes, hence phenotypes.
We will work through the following problem as an example:
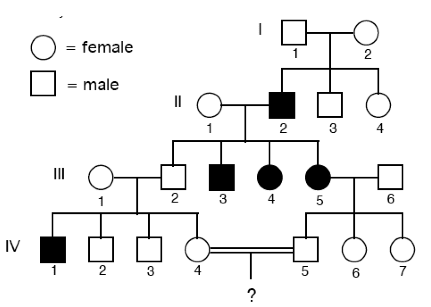
In this pedigree chart, the disease is homozygous recessive. (What does this mean?- Two recessive alleles have to be present in order for the person to display the diseased phenotype).
The question is: What is the probability that the bottom 2 people (4 and 5) have a child with the disease?
Some NCEA questions may require you to determine the phenotype of offspring through a pedigree chart.
We can do this by deducing parent genotypes and then working out possible offspring genotypes, hence phenotypes.
We will work through the following problem as an example:
In this pedigree chart, the disease is homozygous recessive. (What does this mean?- Two recessive alleles have to be present in order for the person to display the diseased phenotype).
The question is: What is the probability that the bottom 2 people (4 and 5) have a child with the disease?
Starting with the left hand side of the diagram, what can we work out?
Now if we look at the right hand side of the diagram.
This gives two possible Punnett squares to be examined:
Punnett square 1: IV5 is carrier (as pre-determined) and IV4 is homozygous dominant
- III:2 is definitely a carrier (Tt) as one parent (II:2) is affected (tt).
- III:1 is also definitely a carrier (Tt) as when mating with III:2 they produce an affected (tt) offspring (IV:1)
- This means that we can work out the possibilities for IV:4 as we know the parent genotypes. It follows the standard arrangement for two carrier parents giving the options of:
- TT (1/4)
- Tt (2/4 = 1/2)
- tt (Normally 1/4 but in this case 0 as individual not marked as affected).
- Therefore for this scenario, the probabilities for IV:4 are :
- TT (1/3)
- Tt (2/3)
Now if we look at the right hand side of the diagram.
- IV:5 is definitely a carrier (Tt) as one of their parents (III:5) is affected.
This gives two possible Punnett squares to be examined:
Punnett square 1: IV5 is carrier (as pre-determined) and IV4 is homozygous dominant
This gives nil affected offspring so we can disregard this option for your question (as we are ONLY looking for scenarios which produce affected individuals).
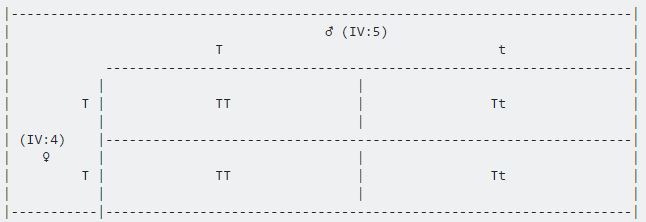
Therefore the alternative Punnett Square 2 is: IV5 is carrier (as pre-determined) and IV4 is heterozygous
Therefore the alternative Punnett Square 2 is: IV5 is carrier (as pre-determined) and IV4 is heterozygous
Giving 1/4 affected offspring.
As mentioned above, in order to have affected offspring then IV:4 must be Tt. There is a 2/3 chance of this being the case. If this is the case, then there is a 1/4 chance of the child being tt.
Both conditions need to be true for this to happen so we multiply the fractions:
2/3 * 1/4 = 1/6
Our answer to the question: What is the probability that the bottom 2 people (IV4 and IV5) have a child with the disease?
--> The probability that IV4 and IV5 have a child with the disease is 1/6.
As mentioned above, in order to have affected offspring then IV:4 must be Tt. There is a 2/3 chance of this being the case. If this is the case, then there is a 1/4 chance of the child being tt.
Both conditions need to be true for this to happen so we multiply the fractions:
2/3 * 1/4 = 1/6
Our answer to the question: What is the probability that the bottom 2 people (IV4 and IV5) have a child with the disease?
--> The probability that IV4 and IV5 have a child with the disease is 1/6.
IDEA CAPSULE 6- Test crosses
What is a test cross and why do we use them?
An individual that displays a dominant trait could be either homozygous dominant (i.e. BB) or heterozygous (i.e. Bb).
In breeding programs, individuals with homozygous genotypes are needed because they are pure breeding and will pass on alleles for the desired trait 100% of the time.
We cannot tell by looking whether an individual we want in our breeding program is homozygous dominant or heterozygous because they both express the dominant phenotype (physical appearance). Therefore, to determine the individuals genotype (hence suitability for breeding) we need to carry out a test cross.
The Test Cross
In a test cross, the individual being tested is crossed with an individual that has the recessive genotype (i.e. bb), hence the recessive trait.
The diagram below, illustrates the two different outcomes of a test cross.
What is a test cross and why do we use them?
An individual that displays a dominant trait could be either homozygous dominant (i.e. BB) or heterozygous (i.e. Bb).
In breeding programs, individuals with homozygous genotypes are needed because they are pure breeding and will pass on alleles for the desired trait 100% of the time.
We cannot tell by looking whether an individual we want in our breeding program is homozygous dominant or heterozygous because they both express the dominant phenotype (physical appearance). Therefore, to determine the individuals genotype (hence suitability for breeding) we need to carry out a test cross.
The Test Cross
In a test cross, the individual being tested is crossed with an individual that has the recessive genotype (i.e. bb), hence the recessive trait.
- The presence of offspring with the recessive trait indicates that the individual being tested is heterozygous (therefore not a pure breeder). It would not be used in the breeding program.
- However if no offspring have the recessive trait after several crossings (to get a large number of offspring/ sample size), then the individual being tested may be assumed to be homozygous and so may be used in the breeding program,
The diagram below, illustrates the two different outcomes of a test cross.
IDEA CAPSULE 7- Complete, incomplete and co-dominance

Complete dominance
Complete dominance occurs when the information on one allele (the dominant allele) is always expressed in the phenotype (physical appearance) when that allele is present in the genotype (genetic makeup) i.e. the presence of a dominant allele masks the presence of a recessive allele.
A recessive allele will only be expressed in the phenotype when both alleles in the genotype are recessive e.g. aa
When both alleles in a genotype are the same e.g. AA or aa, the organism is considered homozygous and a pure breeder because it can only pass on one type of allele to its offspring. However when both alleles are different e.g. Aa, the individual is heterozygous for the trait (and is not a pure breeder because it can pass on either type of allele to its offspring).
Example of complete dominance: Tongue rolling is dominant. When the dominant allele for tongue rolling T is present, the recessive allele t for non-tongue allele is masked in the phenotype and the individual will express the tongue-rolling trait.
Complete dominance occurs when the information on one allele (the dominant allele) is always expressed in the phenotype (physical appearance) when that allele is present in the genotype (genetic makeup) i.e. the presence of a dominant allele masks the presence of a recessive allele.
A recessive allele will only be expressed in the phenotype when both alleles in the genotype are recessive e.g. aa
When both alleles in a genotype are the same e.g. AA or aa, the organism is considered homozygous and a pure breeder because it can only pass on one type of allele to its offspring. However when both alleles are different e.g. Aa, the individual is heterozygous for the trait (and is not a pure breeder because it can pass on either type of allele to its offspring).
Example of complete dominance: Tongue rolling is dominant. When the dominant allele for tongue rolling T is present, the recessive allele t for non-tongue allele is masked in the phenotype and the individual will express the tongue-rolling trait.

Incomplete dominance
This is a pattern of gene expression in which the phenotype of a heterozygous individual is intermediate between those of the parents. It occurs when one allele doesn’t completely dominate over the other. This means that when both alleles are present in the heterozygous genotype, they both contribute to produce a phenotype that is an intermediate or blend of the genetic information. Therefore three different phenotypes may be expressed (instead of only two in complete dominance where the presence of the dominant allele masked the presence of a recessive allele); one for homozygous dominant, another for homozygous recessive and one for a heterozygous organism.
The alleles are hence represented by two of the same capital letters, however one will have a ‘ next to it.
Example of incomplete dominance: Pink Snapdragon flowers (RR’) are often the result of incomplete dominance. When red roses (RR), which contain the dominant red allele (R), are mated with white roses (R’R’), which is recessive (R’), the offspring will be heterozygous (RR’) and express a pink phenotype because the red allele does not completely mask the white allele.
This is a pattern of gene expression in which the phenotype of a heterozygous individual is intermediate between those of the parents. It occurs when one allele doesn’t completely dominate over the other. This means that when both alleles are present in the heterozygous genotype, they both contribute to produce a phenotype that is an intermediate or blend of the genetic information. Therefore three different phenotypes may be expressed (instead of only two in complete dominance where the presence of the dominant allele masked the presence of a recessive allele); one for homozygous dominant, another for homozygous recessive and one for a heterozygous organism.
The alleles are hence represented by two of the same capital letters, however one will have a ‘ next to it.
Example of incomplete dominance: Pink Snapdragon flowers (RR’) are often the result of incomplete dominance. When red roses (RR), which contain the dominant red allele (R), are mated with white roses (R’R’), which is recessive (R’), the offspring will be heterozygous (RR’) and express a pink phenotype because the red allele does not completely mask the white allele.

Co-dominance
Occurs when both alleles are equally dominant. This means when both alleles are present in the heterozygous genotype they are both expressed in the phenotype and as in incomplete dominance, three different phenotypes can occur. However in codominance, the "recessive" & "dominant" traits appear together in the phenotype of hybrid organisms. In codominance, the alleles for the gene are both given capital letters to indicate that they are both dominant alleles.
Example 1 of co-dominance: Roan coat colour in cattle- R for red and W for the white gene for coat colour. The heterozygous offspring RW is a mixture of red and white hairs to give an overall blotchy red and white coat (roan phenotype)
Example 2 of co-dominance: Checkered chicken. The intermediate phenotype (for chickens with a heterozygous genotype) is checkered coat (both black and white feathers are expressed in the intermediate coat).
Occurs when both alleles are equally dominant. This means when both alleles are present in the heterozygous genotype they are both expressed in the phenotype and as in incomplete dominance, three different phenotypes can occur. However in codominance, the "recessive" & "dominant" traits appear together in the phenotype of hybrid organisms. In codominance, the alleles for the gene are both given capital letters to indicate that they are both dominant alleles.
Example 1 of co-dominance: Roan coat colour in cattle- R for red and W for the white gene for coat colour. The heterozygous offspring RW is a mixture of red and white hairs to give an overall blotchy red and white coat (roan phenotype)
Example 2 of co-dominance: Checkered chicken. The intermediate phenotype (for chickens with a heterozygous genotype) is checkered coat (both black and white feathers are expressed in the intermediate coat).
IDEA CAPSULE 8- Multiple and Lethal alleles
|
Multiple alleles
Some genes have more than two different alleles that code for a trait i.e. they have multiple alleles. However, like always, an individual will only have two alleles in its genotype. It is quite common for more than two alleles to control a trait in a population. Multiple alleles can only be studied in a population- as each individual only has two alleles. Example of multiple alleles: Human blood types |
|
Lethal alleles
A lethal allele occurs when a mutation results in an allele that produces a non-functional version of the essential protein. If an individual inherits a lethal combination of alleles it will die shortly after birth.
Lethal alleles may be recessive or dominant depending on the gene or genes involved.
“Carrier”- is the term used for an individual who is heterozygous for a lethal allele or any allele causing a genetic disease.
Example of lethal alleles: Drosophila wings- a mutated allele is dominant and produces curly wings. Fruit flies that inherit a homozygous dominant genotype (CC) for the curly-winged genotype do not survive to hatch from their egg. Offspring with a single mutated allele (i.e. a heterozygous genotype Cc) will not die but will have curly wings.
Human Example of lethal alleles: Brachydactyly is a genetic condition in which the fingers are abnormally short in heterozygotes; however, this condition is fatal during infancy to homozygous recessive individuals due to major skeletal defects.
e.g. Marriage between Two Brachydactyl People:
B b
B BB Bb
b Bb bb
F1 Genotypic Results: 1 BB : 2 Bb : 1 bb
F1 Phenotypic Results: 1 Normal Child : 2 Brachydactyl Children : 1 Infant Death
A lethal allele occurs when a mutation results in an allele that produces a non-functional version of the essential protein. If an individual inherits a lethal combination of alleles it will die shortly after birth.
Lethal alleles may be recessive or dominant depending on the gene or genes involved.
- Dominant Lethal Allele - Quickly eliminated from the population, because usually causes death before the individual can reproduce.
- Recessive Lethal Allele - Will only cause death in the homozygous recessive condition.
“Carrier”- is the term used for an individual who is heterozygous for a lethal allele or any allele causing a genetic disease.
- These individuals are able to live and reproduce.
- Carriers do not phenotypically express the genetic condition.
- These individuals, however, can pass the lethal allele onto offspring.
Example of lethal alleles: Drosophila wings- a mutated allele is dominant and produces curly wings. Fruit flies that inherit a homozygous dominant genotype (CC) for the curly-winged genotype do not survive to hatch from their egg. Offspring with a single mutated allele (i.e. a heterozygous genotype Cc) will not die but will have curly wings.
Human Example of lethal alleles: Brachydactyly is a genetic condition in which the fingers are abnormally short in heterozygotes; however, this condition is fatal during infancy to homozygous recessive individuals due to major skeletal defects.
e.g. Marriage between Two Brachydactyl People:
B b
B BB Bb
b Bb bb
F1 Genotypic Results: 1 BB : 2 Bb : 1 bb
F1 Phenotypic Results: 1 Normal Child : 2 Brachydactyl Children : 1 Infant Death
IDEA CAPSULE 9- Dihybrid inheritance
We've just learned about monohybrid inheritance in IDEA CAPSULE 3 & 4. Recall that monohybrid inheritance is the inheritance of one characteristic controlled by one gene which can have two or more different alleles at the same locus on a homologous pair of chromosomes.
"di" means "two" in greek, hence dihybrid inheritance is looking at the inheritance of two different characteristics, each controlled by their own gene.
There are two main scenarios when it comes to dihybrid inheritance.
Dihybrid inheritance without linkage
Genes that are not linked are found on two separate chromosomes and are therefore segregate independently (during meiosis).
Dihybrid inheritance with linkage
Linked genes occur on the same chromosome, therefore, tend to be inherited together (i.e., do not segregate independently). Linkage actually alters the inheritance pattern.
There are two outcomes that can result from linkage depending on how close/ far apart the genes are located on the chromosome:
Linkage and crossing over have NO EFFECT on the inheritance pattern if the genotypes are HOMOZYGOUS
e.g. If both parents are AABB then all offspring will be AABB.
Linkage and crossing over AFFECT the inheritance pattern if the genotypes are HETEROZYGOUS e.g. AaBb.
The following diagram shows dihybrid inheritance without linkage (right side) and with linkage (left side)
We've just learned about monohybrid inheritance in IDEA CAPSULE 3 & 4. Recall that monohybrid inheritance is the inheritance of one characteristic controlled by one gene which can have two or more different alleles at the same locus on a homologous pair of chromosomes.
"di" means "two" in greek, hence dihybrid inheritance is looking at the inheritance of two different characteristics, each controlled by their own gene.
There are two main scenarios when it comes to dihybrid inheritance.
- Dihybrid inheritance without linkage
- Dihybrid inheritance with linkage
Dihybrid inheritance without linkage
Genes that are not linked are found on two separate chromosomes and are therefore segregate independently (during meiosis).
Dihybrid inheritance with linkage
Linked genes occur on the same chromosome, therefore, tend to be inherited together (i.e., do not segregate independently). Linkage actually alters the inheritance pattern.
There are two outcomes that can result from linkage depending on how close/ far apart the genes are located on the chromosome:
- When two linked genes are close together on the chromosome, crossing over during recombination in meiosis is unlikely to occur
- If two genes are far apart on the chromosome, crossing over is likely to occur.
Linkage and crossing over have NO EFFECT on the inheritance pattern if the genotypes are HOMOZYGOUS
e.g. If both parents are AABB then all offspring will be AABB.
Linkage and crossing over AFFECT the inheritance pattern if the genotypes are HETEROZYGOUS e.g. AaBb.
The following diagram shows dihybrid inheritance without linkage (right side) and with linkage (left side)
As we can see in the left diagram, crossing over occurs between the non-sister chromatids (during Prophase I of meiosis) so that linked alleles are crossed over to give new combinations, Ab and aB (these are called recombinants). The parental combinations for the genotypes of these gametes are AB and ab while the recombinants and the genotypes of these gametes is Ab and aB.
How do we know if two genes are linked or not?
The standard method to see the effect of crossing over is to compare the results of a test cross without linkage of the genes and a test cross with linkage of the genes.
Under no linkage:
AaBb x aabb
The expected phenotype ratio of the offspring is 1 tall yellow: 1 tall green: 1 short yellow: 1 short green
However when genes are linked:
AaBb x aabb
Most gametes from the AaBb parent are AB and ab, however there are a few recombinants (Ab and aB).
The F1 offspring no longer display the 1: 1: 1: 1 ratio of phenotypes.
With linkage and crossing over, the F1 phenotypes are mostly the parental types (tall yellow and short green) at an approx. 1:1 ratio, however a few are the recombinant phenotypes, tall green, short yellow.
A different explanation of linkage....
Here is a link to a different website which may explain the concept of linked genes better (it has pictures!)- Scroll down to the heading "10.2.5: Dihybrid Cross with Linkage".
How do we know if two genes are linked or not?
The standard method to see the effect of crossing over is to compare the results of a test cross without linkage of the genes and a test cross with linkage of the genes.
Under no linkage:
AaBb x aabb
The expected phenotype ratio of the offspring is 1 tall yellow: 1 tall green: 1 short yellow: 1 short green
However when genes are linked:
AaBb x aabb
Most gametes from the AaBb parent are AB and ab, however there are a few recombinants (Ab and aB).
The F1 offspring no longer display the 1: 1: 1: 1 ratio of phenotypes.
With linkage and crossing over, the F1 phenotypes are mostly the parental types (tall yellow and short green) at an approx. 1:1 ratio, however a few are the recombinant phenotypes, tall green, short yellow.
A different explanation of linkage....
Here is a link to a different website which may explain the concept of linked genes better (it has pictures!)- Scroll down to the heading "10.2.5: Dihybrid Cross with Linkage".
IDEA CAPSULE 10- Punnett Squares for dihybrid crosses
Use the powerpoint above as a guide for how to use a Punnett Square for dihybrid crosses. (It works through an example involving Mendel's Pea plants).
Steps to remember:
- Find out what your parent genotypes are and write them down e.g. AaBB and aabb
- Determine what the different combinations of alleles could be in the gametes of each parent. You can use the technique "FOIL" to work out all possible combinations. FOIL stands for "Firsts" AaBB, "Outsides" AaBB, "Insides" AaBB, Lasts, AaBB. In this example we can see that for parent AaBB, the gametes can either have a combination of alleles AB or aB.
- Write the different combinations for the female parent in the horizontal row and the male parent in the vertical rows.
- Determine the final genotypes of the offspring, just like we did in monohybrid crosses, however this time, the offspring genotypes will consist of 4 alleles.
- REMEMBER: When writing the offspring genotypes, keep alleles from the same trait together and always put a dominant allele first if present e.g. AaBB (correct) NOT aBAB (alleles from the same trait not together, recessive allele in front of dominant allele).
- We can then work out the genotype ratios and phenotype ratios of the offspring as usual.
Try these dihybrid cross examples in the file below (Spongebob Dihybrid Cross Worksheet)
- Print off the worksheet and show your working to how you get your answers (if required) e.g. draw Punnett Squares of necessary
- When you're finished, show me your answers and I can link you the answers so you can self-mark.
| Spongebob Dihybrid Cross Worksheet | |
| File Size: | 176 kb |
| File Type: | |
IDEA CAPSULE 11- Intoduction to Genetic Variation in Populations
This 5 fingers of evolution video and additional worksheet will introduce you to some of the concepts we will be looking at next. In this section we look at how genetic variation arises/ can change within populations of individuals.
- Watch the video once through first
- Then see if you can fill in the worksheet
- If you are having trouble, you can always go back to the video to find your answers
|
|
|
IDEA CAPSULE 12- Types of selection
There are three main types of selection which can act on a population
The following powepoint will give you an insight into what these selection pressures are and how they can affect genetic populations.
There are three main types of selection which can act on a population
- Natural Selection
- Sexual Selection
- Artificial Selection
The following powepoint will give you an insight into what these selection pressures are and how they can affect genetic populations.
Associated activity: Try and find some other real-life examples of natural selection, sexual selection and artificial selection. You may refer to webpages, videos, graphics or books to do this.
IDEA CAPSULE 13- Factors that increase genetic variation in a pop.
There are several factors which increase genetic variation in a population:
1. Random mating, random fertilization, and recombination between homologous chromosomes during meiosis.
(Refer back to IDEA CAPSULE 2 for a better explanation of this).
2. Mutations
Mutations are one of the ways that new alleles can appear in a gene pool, therefore mutation increases genetic variation in a population and is essential for evolution.
Natural selection will act on the mutation.
Mutations will only enter the gene pool and become subject to natural selection if the mutation occurs during meiosis. This is because mutations can only be inherited by offspring if they occur in the reproductive cells (gametes). Mutations that occur during DNA replication for mitosis in the somatic (body) cells of organisms cannot be inherited, therefore do not enter the gene pool.
There are 2 different categories of mutations:
Note: We learn a lot more about how these mutations arise in the standard "Biology 2.7- Gene Expression".
There are several factors which increase genetic variation in a population:
1. Random mating, random fertilization, and recombination between homologous chromosomes during meiosis.
(Refer back to IDEA CAPSULE 2 for a better explanation of this).
2. Mutations
Mutations are one of the ways that new alleles can appear in a gene pool, therefore mutation increases genetic variation in a population and is essential for evolution.
Natural selection will act on the mutation.
- If it is favourable, the organism will be more likely to reproduce and pass on the allele to its offspring and the frequency of the mutated allele will increase.
- However if it is unfavourable, the organism is less likely to reproduce, hence the frequency of the allele is likely to decrease or be diminished in the population.
Mutations will only enter the gene pool and become subject to natural selection if the mutation occurs during meiosis. This is because mutations can only be inherited by offspring if they occur in the reproductive cells (gametes). Mutations that occur during DNA replication for mitosis in the somatic (body) cells of organisms cannot be inherited, therefore do not enter the gene pool.
There are 2 different categories of mutations:
- Neutral/ silent mutations- neither favourable nor unfavourable so they are not subject to selection. We get these mutations when a base change in a DNA molecule may not produce a different amino acid. This occurs due to redundancy in the genetic code (a single amino acid may be coded for by more than one codon).
- Favourable/ unfavourable mutations- will be selected for or against.
Note: We learn a lot more about how these mutations arise in the standard "Biology 2.7- Gene Expression".
|
3. Movement- Immigration
Migration is the movement of individuals from one population to another. Migration into or out of a population may be responsible for a marked change in allele frequencies (the proportion of members carrying a particular variant of a gene). In population genetics, gene flow (also known as gene migration) is the transfer of alleles or genes from one population to another. The smaller the population, the more likely migration will have a significant effect on allele frequency. However, if the population is large, migration may have little or no effect on allele frequency. --> Immigration involves individuals migrating INTO a population. Immigration can add new alleles to a population, therefore increasing a population’s genetic biodiversity. Associated activity: Modelling migration and mutations, an active scenario (Note: We will do this in class together) --> |
|
IDEA CAPSULE 14- Factors that decrease genetic variation in a pop.
There are several factors which decrease genetic variation in a population:
1. All types of Selection (Sexual, Natural and Artificial)
Selection pressures reduce genetic variation in a population because they select for the most favourable traits only.
For example, natural selection is the process whereby organisms better adapted to their environment tend to survive and produce more offspring. Thus, genetic variation in a population is usually decreased as only organisms with the most favourable traits are able to survive and reproduce.
Remember, natural selection is the main process which can lead to evolution/
2. Movement- Emigration
As you will recall, migration is the movement of individuals from one population to another.
--> Emigration involves individuals migrating OUT of a population. Emigration could remove alleles from a population, hence reducing a population’s genetic biodiversity.
3. Genetic Drift
The change in allele frequencies due to chance.
Genetic drift is most likely to have an effect on small populations. Because when populations are large, and mating is random, allele frequencies tend to remain stable from generation to generation (unless natural selection is operating). However, when populations are small it is likely that allele frequencies will change from generation to generation by chance.
Think about it:
What is the chance of losing a recessive allele “a” if there is a population of two, heterozygous organisms and they have two children?
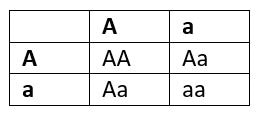
Aa x Aa
Only chance of losing “a” allele if the offspring has the genotype AA. Therefore ¼ chance that the “a” allele will be lost if the couple has one child. However, the more children they have, the lower the chance of losing the “a” allele. The chance of losing the “a” allele when having two children is ¼ x ¼= 1/16.
There are several factors which decrease genetic variation in a population:
1. All types of Selection (Sexual, Natural and Artificial)
Selection pressures reduce genetic variation in a population because they select for the most favourable traits only.
For example, natural selection is the process whereby organisms better adapted to their environment tend to survive and produce more offspring. Thus, genetic variation in a population is usually decreased as only organisms with the most favourable traits are able to survive and reproduce.
Remember, natural selection is the main process which can lead to evolution/
2. Movement- Emigration
As you will recall, migration is the movement of individuals from one population to another.
--> Emigration involves individuals migrating OUT of a population. Emigration could remove alleles from a population, hence reducing a population’s genetic biodiversity.
3. Genetic Drift
The change in allele frequencies due to chance.
Genetic drift is most likely to have an effect on small populations. Because when populations are large, and mating is random, allele frequencies tend to remain stable from generation to generation (unless natural selection is operating). However, when populations are small it is likely that allele frequencies will change from generation to generation by chance.
Think about it:
What is the chance of losing a recessive allele “a” if there is a population of two, heterozygous organisms and they have two children?
Aa x Aa
Only chance of losing “a” allele if the offspring has the genotype AA. Therefore ¼ chance that the “a” allele will be lost if the couple has one child. However, the more children they have, the lower the chance of losing the “a” allele. The chance of losing the “a” allele when having two children is ¼ x ¼= 1/16.
|
4. Founders Effect
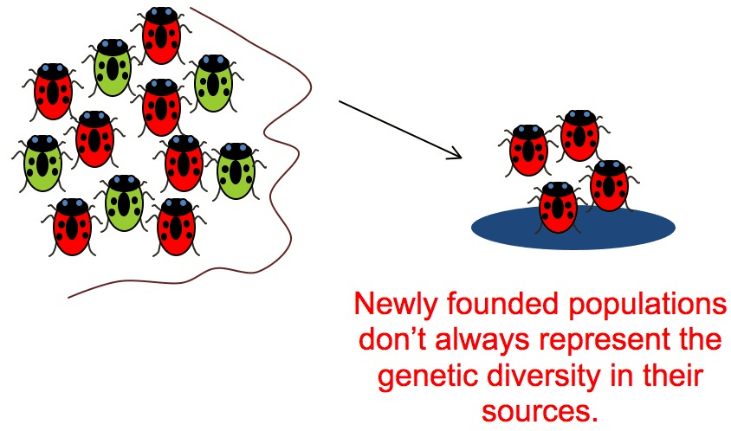
A small group of individuals that colonises a new isolated area (such as an island) is called a founder population. Founder populations often form because a small group of individuals becomes separated from the larger, original population (likely due to an environmental cause e.g. high winds, flooding or water currents). The range and frequency of the alleles present in this small (founder) group is unlikely to be representative of the original population e.g. some alleles may not be present at all and others may be less frequent or more frequent. Effects on a founder population: Because a founder population is small, it is likely to:
|
|
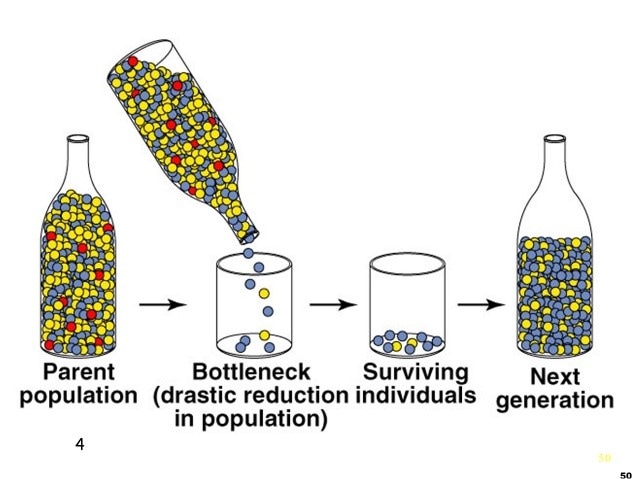
5. Bottleneck Effect
The bottleneck effect occurs when a population is suddenly reduced in numbers to a small size. This typically occurs in one of two ways:
As a result of population numbers falling, it is likely that the range of alleles in the bottle neck population will decrease and the frequencies of alleles in the resulting population will change. Even if the population increases again to its original size or becomes larger, it will still have reduced genetic biodiversity. |
|
|
Associated activity: Read the famous New Zealand example of the bottle-neck effect by clicking on the file below. What other famous examples can you find where populations have been reduced to a bottleneck and then recovered? Has this had any effect on the populations wellbeing?
| |||||||
ACTIVITIES (QUIZZES, GAMES, QUESTIONS AND MORE)
|
ACTIVITY 1
|
|
ACTIVITY 2
|
|
ACTIVITY 3
|
|
ACTIVITY 4
|
VIDEO RESOURCES
Useful videos from some of our favourite content producers:
TEST YOURSELF (THINK YOU'VE GOT IT ALL FIGURED OUT? HAVE A GO AT SOME OF THESE!)
- Have a go at answering the questions on THIS site